Cell Screening
Nikon's incubator-style imaging systems elevate live cell imaging to a new level of environmental control. Sophisticated temperature, humidity and gas control combine with automated plate management and advanced imaging tools to provide the ultimate imaging system for long-term monitoring of delicate samples.
Product Lineup
New
ECLIPSE Ji - Digital Microscopes
The new ECLIPSE Ji Digital Inverted Microscope provides research microscope power in a benchtop assay instrument.
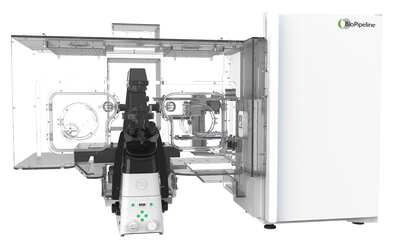
BioPipeline LIVE - High Content Imaging
Environmentally-controlled stable long-term imaging and analysis of multiple samples in a range of imaging modalities.